Resources

notame
notame is an R package co-developed at the University of Turku, University of Eastern Finland, and Afekta Technologies for nontargeted metabolomics data processing. It features a streamlined workflow consisting of reading in exported data from signal detection software (currently MS-DIAL), correcting signal intensity drift and batch effects, identifying and flagging low-quality molecular features, imputing missing values, correcting batch effects, and clustering similar molecular features. notameStats is a separate package in the bundle that includes a variety of useful statistical tests for metabolomics data, and notameViz provides an array of data visualization features. We currently use these tools in all our metabolomics projects involving medium to large sample sets.
Want to know more? Visit the GitHub or Bioconductor links below to install and get started with the packages, or check out a book chapter on the practical use of the packages, available as a preprint!
Install the latest Bioconductor versions of notame, notameStats, and notameViz in R:
if (!require(“BiocManager”, quietly = TRUE))
install.packages(“BiocManager”)
BiocManager::install(“notame”)
BiocManager::install(“notameStats”)
BiocManager::install(“notameViz”)Citation:
Klåvus, A., Kokla, M., Noerman, S., Koistinen, V. M., Tuomainen, M., Zarei, I., ... & Hanhineva, K. (2020). “Notame”: workflow for non-targeted LC–MS metabolic profiling. Metabolites, 10(4), 135.Monoisotopic mass calculator
This is a simple calculator for the monoisotopic masses of any neutral molecule and its most common adducts in LC-MS. The formula needs to be given without brackets. The element masses used in this calculator are those of the most abundant isotopes (thus giving only single-isotope values), and they are based on the CODATA Internationally recommended 2022 values of the Fundamental Physical Constants, in unified atomic mass units (u / Da) rounded to six decimals. Carbon-12, which has six protons and six neutrons, is defined as exactly 12 u (or 12 Da). The monoisotopic mass of a hydrogen atom is used for the neutral monoisotopic mass, whereas the mass of a proton (without the electron) is used for calculating the adduct masses.
Mass error calculator
Our mass error calculator will calculate the mass error in millidaltons (mDa) and parts per million (ppm) between an observed m/z and the theoretical m/z calculated based on the molecular formula of the annotated metabolite and the same adducts as in the monoisotopic mass calculator.
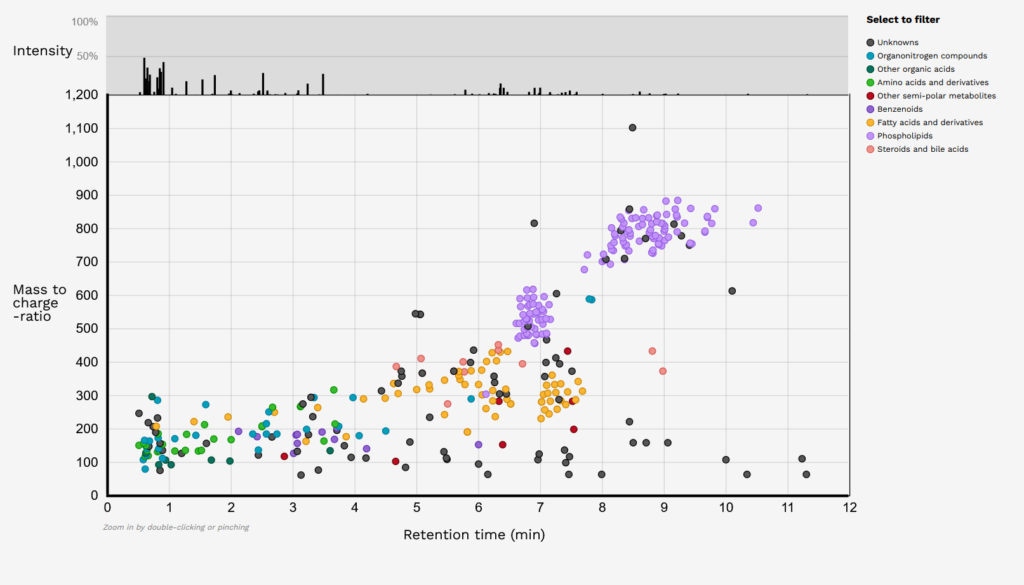
Plasma Benchmark Metabolite Visualisation Application
Plasma benchmark is a collaboration project between the Chalmers University of Technology, International Agency for Research on Cancer (IARC) in Lyon, France, the University of Eastern Finland, and the University of Turku. This project aims to robustly determine and catalogue the key human plasma metabolites and lipids detectable with typical LC-MS metabolomics approaches and to create a benchmark for a more comprehensive and reproducible way to assess the plasma metabolome in clinical human studies. We have designed an interactive platform to view the project results.
We are currently expanding the database and many typical plasma metabolites may still be missing. This is because we require that the signal of the metabolite must be reproducibly detected in the participating laboratories, which use UPLC-QTOF-MS instruments and a method modified from our untargeted metabolomics approach (Klåvus et al. 2020).
Visit our GitHub page for more resources, such as:
Outreach material including posters and presentations →
